From genetics to computational biology: a journey turning data into healthcare insights
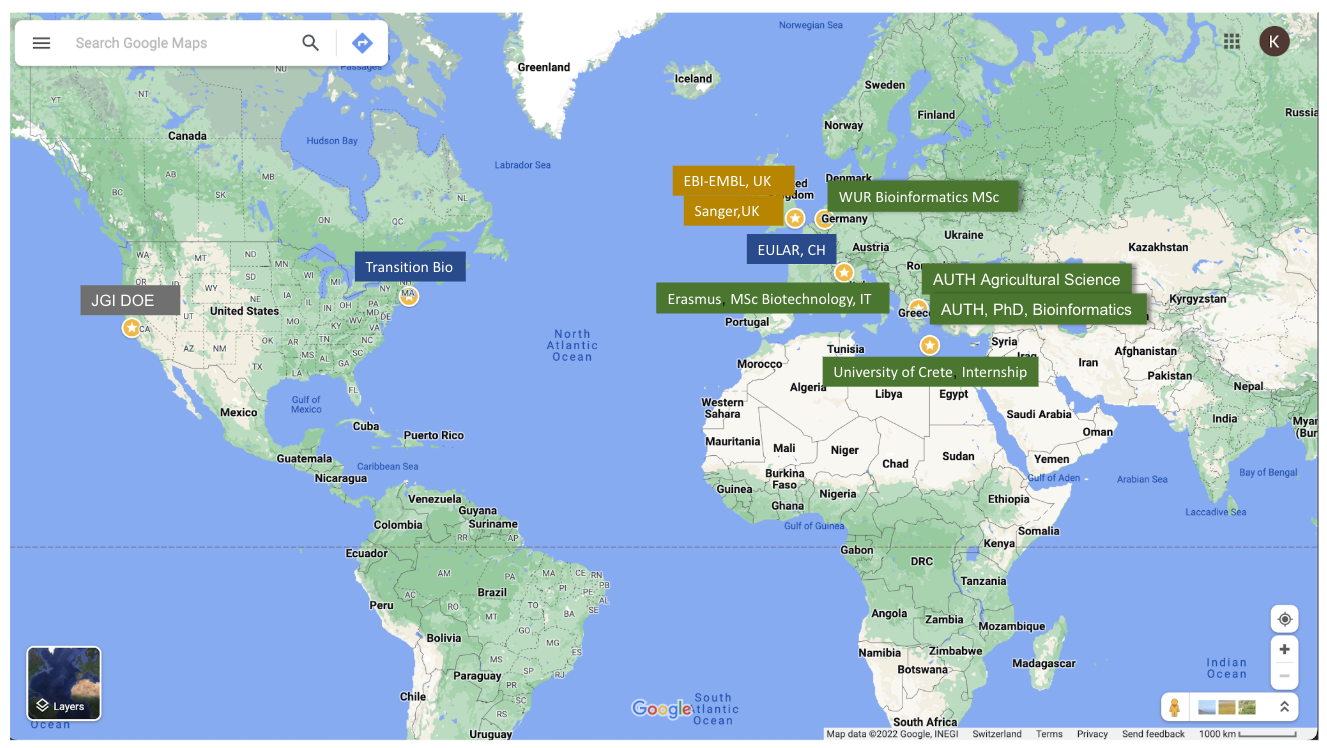
First Studies
My research career bridges genetics, bioinformatics, and data analysis, guided by a commitment to turning data into insights that advance both scientific discovery and healthcare applications. I began with a strong foundation in genetics and plant biology at Aristotle University of Thessaloniki, complemented by a six-month master’s-level exchange in biotechnology at Università degli Studi di Milano – Facoltà di Agraria. I then pursued a Master’s in Bioinformatics at Wageningen University, where my thesis explored transcription factors and secreted proteins in host–pathogen interactions. An internship at the Institute of Molecular Biology and Biotechnology (IMBB-FORTH) in Crete introduced me to plant RNA interference (RNAi), strengthening both my wet-lab and computational skills and cementing my transition into computational biology.
Advancing Computational Biology, RNA-seq and PhD at JGI
My professional work began in 2010 at the Joint Genome Institute (JGI) under the U.S. Department of Energy focused on developing workflows and visualization tools for early RNA sequencing (RNA-seq) - Early RNA-seq pipeline - data analysis, essential for microbial research. At a time when RNA-seq was an emerging technique, I developed one of the first pipelines for expression data and web tools within the Integrated Microbial Genomes (IMG) system.
This role sparked my PhD research, examining transcriptomic responses to environmental stress in bacteria. My thesis revealed adaptive mechanisms in response to stress, including:
- Differential gene expression,
- Evidence of antisense genes,
- Transcriptional mutagenesis and variants,
- Reverse transcriptase–like protein.
My PhD was conducted later while working full-time for European Bioinformatics Institute - EMBL and in collaboration with Aristotle University, National Energy Research Scientific Computing Center and JGI.
Pioneering Genomic Resources with Ensembl
A defining stage of my career began in 2013 when I joined Ensembl at the European Bioinformatics Institute (EMBL-EBI) and the Wellcome Sanger Institute. Here, I contributed to one of the world’s most critical genomic resources: the Ensembl platform. Key projects included:
- GRCh38 human genome annotation (the current human reference genome)
- Illumina Human BodyMap project
- The Human Pangenome Project (featured on the cover of Nature)
- Pipelines for non-coding RNAs (lncRNA) and immune genes (IG/TR)
I also led the first multigenome annotations (including primates and rodents), helping establish the foundation for scalable, high-throughput genome analysis at Ensembl. These efforts contributed to Ensembl’s role in global genetics by delivering high-quality genomic data to researchers, clinicians, and pharmaceutical R&D teams. In parallel, I supervised and mentored data scientists, introducing bioinformatics best practices, cloud infrastructure, and containerized workflows into production pipelines.
International Collaborations and Industry Partnerships
During my time at Ensembl, I collaborated closely with academic groups and industry partners adapting genomic data resources to diverse research and translational needs. I contributed to 20+ custom genomics projects, spanning:
- Model organism annotations
- Single-cell transcriptomics
- Biomarker discovery
- Therapeutic target identification
- Transmissible cancer studies
- Population studies
Translational Bioinformatics and Healthcare Data
Following Ensembl, I transitioned into translational bioinformatics roles bridging research, healthcare and pharma events.
At Transition Bio (Cambridge & Boston), I integrated multi-omics data and machine learning predictions into drug discovery, focusing on biomolecular condensates and therapeutic targets.
At EULAR – European Alliance of Associations for Rheumatology, I built secure cloud-based platforms for clinical data collection and omics data analysis, enabling:
- Direct data collection from patients
- Data visualization for advocacy
- Bulk RNA-seq, scRNA-seq, BCR/TCR-seq analysis
These projects required close collaboration with wet-lab scientists, patient groups, clinicians, advocacy organizations, and pharmaceutical partners.
Publications and Research Impact
I have authored 26+ publications in high-impact journals including Nature, Science, Cell Reports Methods, and Nucleic Acids Research, with over 20,700 citations to date.
🔬 Research Highlights
-
Human Pangenome Project
The Nature cover story on the draft human pangenome reference, Ensembl Annotation (Nature, Cell Reports Methods, 2023). -
Comparative Genomics of Unique Species
Explored evolutionary insights from the Tuatara genome (Nature, 2020) and transmissible cancers in Tasmanian devils (Science, 2023). -
Ensembl Genome Annotation
Long-standing contributor to the Ensembl project, with annual updates published in Nucleic Acids Research from 2014 to 2022, including work on farm animal genomes, chicken gene annotation, and lncRNA expression. -
Microbial Transcriptomics & Stress Response
Investigated transcriptomic stress responses in cyanobacteria (PLoS One, 2014) and lignocellulolytic microbial communities (Biotechnology for Biofuels, 2014). -
Structural Variation & Haplotype-Resolved Genomes
Contributed to studies on structural variation in cattle, pigs, and primates (Nature Communications, Gigascience, Mobile DNA). -
Non-Coding RNA & Embryonic Development
Participated in RNAcentral and zebrafish mRNA expression time course projects (NAR, eLife, 2017–2019), and studies on embryonic lethality signatures (Nature Communications, 2019). -
Healthcare & Policy
Recent work includes the EULAR RheumaFacts project on inequity in access to care for rheumatic diseases (Annals of the Rheumatic Diseases, 2025).
Mentorship and Community Engagement
Beyond research, I have a strong commitment to training and outreach.
- Delivered 15+ international workshops (RNA-seq, Ensembl, IMG)
- Tutor in the MSc Host-microbe program at the University of Thessaly
- Google Summer of Code mentor
- Active member of the Hellenic Bioinformatics Association
Research Vision
Looking ahead, my goal is to advance precision medicine and predictive modeling by integrating multi-omics data with bioinformatics pipelines and AI.
Summary
My career embodies a dedication to innovation, collaboration, and impact - spanning foundational genomics projects at Ensembl, international consortia, translational bioinformatics in healthcare, and community engagement worldwide.
